elephant.spade.spade¶
-
elephant.spade.spade(spiketrains, bin_size, winlen, min_spikes=2, min_occ=2, max_spikes=None, max_occ=None, min_neu=1, approx_stab_pars=None, n_surr=0, dither=array(15.0) * ms, spectrum='#', alpha=None, stat_corr='fdr_bh', surr_method='dither_spikes', psr_param=None, output_format='patterns', **surr_kwargs)[source]¶ Perform the SPADE [1], [2], [3] analysis for the parallel input spiketrains. They are discretized with a temporal resolution equal to bin_size in a sliding window of winlen*bin_size.
First, spike patterns are mined from the spiketrains using a technique called frequent itemset mining (FIM) or formal concept analysis (FCA). In this framework, a particular spatio-temporal spike pattern is called a “concept”. It is then possible to compute the stability and the p-value of all pattern candidates. In a final step, concepts are filtered according to a stability threshold and a significance level alpha.
Parameters: - spiketrainslist of neo.SpikeTrain
List containing the parallel spike trains to analyze
- bin_sizepq.Quantity
The time precision used to discretize the spiketrains (binning).
- winlenint
The size (number of bins) of the sliding window used for the analysis. The maximal length of a pattern (delay between first and last spike) is then given by winlen*bin_size
- min_spikesint, optional
Minimum number of spikes of a sequence to be considered a pattern. Default: 2
- min_occint, optional
Minimum number of occurrences of a sequence to be considered as a pattern. Default: 2
- max_spikesint, optional
Maximum number of spikes of a sequence to be considered a pattern. If None no maximal number of spikes is considered. Default: None
- max_occint, optional
Maximum number of occurrences of a sequence to be considered as a pattern. If None, no maximal number of occurrences is considered. Default: None
- min_neuint, optional
Minimum number of neurons in a sequence to considered a pattern. Default: 1
- approx_stab_parsdict or None, optional
Parameter values for approximate stability computation.
- ‘n_subsets’: int
Number of subsets of a concept used to approximate its stability. If n_subsets is 0, it is calculated according to to the formula given in Babin, Kuznetsov (2012), proposition 6:
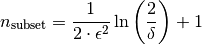
- ‘delta’float, optional
delta: probability with at least
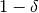
- ‘epsilon’float, optional
epsilon: absolute error
- ‘stability_thresh’None or list of float
List containing the stability thresholds used to filter the concepts. If stability_thresh is None, then the concepts are not filtered. Otherwise, only concepts with
- intensional stability is greater than stability_thresh[0] or
- extensional stability is greater than stability_thresh[1]
are further analyzed.
- n_surrint, optional
Number of surrogates to generate to compute the p-value spectrum. This number should be large (n_surr >= 1000 is recommended for 100 spike trains in spiketrains). If n_surr == 0, then the p-value spectrum is not computed. Default: 0
- ditherpq.Quantity, optional
Amount of spike time dithering for creating the surrogates for filtering the pattern spectrum. A spike at time t is placed randomly within [t-dither, t+dither] (see also
elephant.spike_train_surrogates.dither_spikes()). Default: 15*pq.ms- spectrum{‘#’, ‘3d#’}, optional
Define the signature of the patterns.
- ‘#’: pattern spectrum using the as signature the pair:
(number of spikes, number of occurrences)
- ‘3d#’: pattern spectrum using the as signature the triplets:
(number of spikes, number of occurrence, difference between last and first spike of the pattern)
Default: ‘#’
- alphafloat, optional
The significance level of the hypothesis tests performed. If alpha is None, all the concepts are returned. If 0.<alpha<1., the concepts are filtered according to their signature in the p-value spectrum. Default: None
- stat_corrstr
Method used for multiple testing. See:
test_signature_significance()Default: ‘fdr_bh’- surr_methodstr
Method to generate surrogates. You can use every method defined in
elephant.spike_train_surrogates.surrogates(). Default: ‘dither_spikes’- psr_paramNone or list of int or tuple of int
This list contains parameters used in the pattern spectrum filtering:
- psr_param[0]: correction parameter for subset filtering
(see h_subset_filtering in
pattern_set_reduction()).- psr_param[1]: correction parameter for superset filtering
(see k_superset_filtering in
pattern_set_reduction()).- psr_param[2]: correction parameter for covered-spikes criterion
(see l_covered_spikes in
pattern_set_reduction()).
- output_format{‘concepts’, ‘patterns’}
Distinguish the format of the output (see Returns). Default: ‘patterns’
- surr_kwargs
Keyword arguments that are passed to the surrogate methods.
Returns: - outputdict
- ‘patterns’:
If output_format is ‘patterns’, see the return of
concept_output_to_patterns()If output_format is ‘concepts’, then output[‘patterns’] is a tuple of patterns which in turn are tuples of
- spikes in the pattern
- occurrences of the pattern
For details see
concepts_mining().- if stability is calculated, there are also:
- intensional stability
- extensional stability
For details see
approximate_stability().- ‘pvalue_spectrum’ (only if n_surr > 0):
A list of signatures in tuples format:
- size
- number of occurrences
- duration (only if spectrum==’3d#’)
- p-value
- ‘non_sgnf_sgnt’: list
Non significant signatures of ‘pvalue_spectrum’.
Notes
If detected, this function will use MPI to parallelize the analysis.
References
[1] Torre, E., Picado-Muino, D., Denker, M., Borgelt, C., & Gruen, S. (2013). Statistical evaluation of synchronous spike patterns extracted by frequent item set mining. Frontiers in Computational Neuroscience, 7. [2] Quaglio, P., Yegenoglu, A., Torre, E., Endres, D. M., & Gruen, S. (2017). Detection and Evaluation of Spatio-Temporal Spike Patterns in Massively Parallel Spike Train Data with SPADE. Frontiers in Computational Neuroscience, 11. [3] Stella, A., Quaglio, P., Torre, E., & Gruen, S. (2019). 3d-SPADE: Significance evaluation of spatio-temporal patterns of various temporal extents. Biosystems, 185, 104022. Examples
The following example applies SPADE to spiketrains (list of neo.SpikeTrain).
>>> from elephant.spade import spade >>> import quantities as pq >>> bin_size = 3 * pq.ms # time resolution to discretize the spiketrains >>> winlen = 10 # maximal pattern length in bins (i.e., sliding window) >>> result_spade = spade(spiketrains, bin_size, winlen)
