elephant.spike_train_correlation.covariance¶
-
elephant.spike_train_correlation.covariance(binned_spiketrain, binary=False, fast=True)[source]¶ Calculate the NxN matrix of pairwise covariances between all combinations of N binned spike trains.
For each pair of spike trains
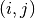 , the covariance
, the covariance ![C[i,j]](../../../_images/math/bf98d44b10da7feeb3ab48513f947f6939e208a9.png) is obtained by binning
is obtained by binning 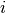 and
and 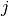 at the desired bin size. Let
at the desired bin size. Let
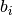 and
and 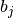 denote the binned spike trains and
denote the binned spike trains and
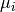 and
and 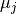 their respective averages. Then
their respective averages. Then![C[i,j] = <b_i-\mu_i, b_j-\mu_j> / (L-1)](../../../_images/math/f4c667d64eaf41a4ea1d13db063306e6e397aa13.png)
where <., .> is the scalar product of two vectors, and
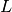 is the
number of bins.
is the
number of bins.For an input of N spike trains, an N x N matrix is returned containing the covariances for each combination of input spike trains.
If binary is True, the binned spike trains are clipped to 0 or 1 before computing the covariance, so that the binned vectors
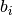 and
and
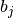 are binary.
are binary.Parameters: - binned_spiketrain(N, ) elephant.conversion.BinnedSpikeTrain
A binned spike train containing the spike trains to be evaluated.
- binarybool, optional
If True, the spikes of a particular spike train falling in the same bin are counted as 1, resulting in binary binned vectors
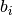 . If
False, the binned vectors
. If
False, the binned vectors 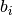 contain the spike counts per bin.
Default: False.
contain the spike counts per bin.
Default: False.- fastbool, optional
If fast=True and the sparsity of binned_spiketrain is > 0.1, use np.cov(). Otherwise, use memory efficient implementation. See Notes [2]. Default: True.
Returns: - C(N, N) np.ndarray
The square matrix of covariances. The element
![C[i,j]=C[j,i]](../../../_images/math/709f60b89db15874f0deced9152a19286b30d59d.png) is
the covariance between binned_spiketrain[i] and
binned_spiketrain[j].
is
the covariance between binned_spiketrain[i] and
binned_spiketrain[j].
Raises: - MemoryError
When using fast=True and binned_spiketrain shape is large.
Warns: - UserWarning
If at least one row in binned_spiketrain is empty (has no spikes).
See also
correlation_coefficient- Pearson correlation coefficient
Notes
- The spike trains in the binned structure are assumed to cover the complete time span [t_start, t_stop) of binned_spiketrain.
- Using fast=True might lead to MemoryError. If it’s the case, switch to fast=False.
Examples
Generate two Poisson spike trains
>>> import neo >>> from quantities import s, Hz, ms >>> from elephant.spike_train_generation import homogeneous_poisson_process >>> from elephant.conversion import BinnedSpikeTrain >>> st1 = homogeneous_poisson_process( ... rate=10.0*Hz, t_start=0.0*s, t_stop=10.0*s) >>> st2 = homogeneous_poisson_process( ... rate=10.0*Hz, t_start=0.0*s, t_stop=10.0*s) >>> cov_matrix = covariance(BinnedSpikeTrain([st1, st2], bin_size=5*ms)) >>> print(cov_matrix[0, 1]) -0.001668334167083546
